Hello,
I’m running CMAQ-ISAM v5.5 built with gcc compilers on a Cray machine, with custom defined regions. I’m using 12US1 domain, and 2022 EMP (v1) emissions. I’ve tested and run ISAM when using states as the emission source regions. However, when using custom defined regions (with files generated using shp2cmaq and defined as RTO_REGIONS below), I see warnings like below:
Could not read RTO_01 from file RTO_REGIONS
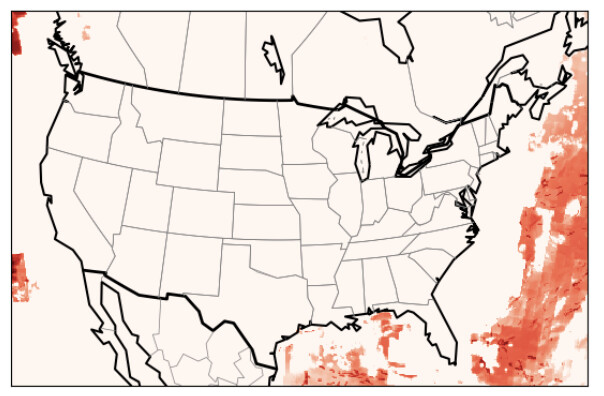
The model, however, continues to run and and produces out files. When examining the output, I see unexpected results - essentially no concentration attributed to the source region defined in RTO_REGIONS. Below I have two screenshots of concentrations produced by two different runs: one from R10 (which covers the state of Texas) on left, and another when using a different ISAM_REGIONS which uses states (in this case TX) - the two runs have same input data except for the region definitions (both maps are for NO2 from corresponding regions, and for same time step: Feb 01 2022 Hour 00):
[Note that I can see impacts from other states as well (used NY, TN, CO as examples) but none from corresponding RTO_REGIONS]
One can see that while impacts of TX emissions are clearly visible, no such thing is seen for R01 (which corresponds to TX). For these simulations, I’ve updated the DESID registry and defined the appropriate tags in the ISAM Control file (screenshots of DESID registry, and the regions netcdf file included below).
I’d really appreciate if someone points to what could be causing an issue here?
A related question is: what is the difference between “Region Label” and “Variable on File” data in the DESID region namelist? My guess is it creates a mapping between the region names in the netCDF region file to the regions used in the ISAM control file. Is there any documentation which explains this more?
Another ISAM specific feedback: some of the jobs (from prior runs, not the ones used for the analysis above) produced error messages indicating out of memory but the model continued to run (w/o producing any outputs) for several hours before running out of walltime - only then I noticed this error message. I’d expect the the job to fail immediately in this case, but I didn’t see that happen.
Best,
Vikram
Based on the error message “Warning in subroutine RDTFLAG - Time step not availabe in file RTO_REGIONS for variable RTO_01”, I am guessing that there is an issue with the time step structure in the RTO_REGIONS file generated by shp2cmaq.
Specifically, these region mask files are expected to be time-independent which should be indicated by a TSTEP attribute of 0 (see section “Time Step Structure” here). I am guessing that the file which leads to realistic results used a TSTEP=0 attribute while the file that leads to unrealistic results doesn’t.
I’m tagging @barronh for further thoughts on the behavior of shp2cmaq and @sergey on further thoughts about how ISAM calculates source tag contributions when a tag is defined with a region but the reading of the variable defining that region from the gridmask file fails as in your case.
Yes, your understanding is correct. You could also just use
ALL', 'RTO_REGIONS', 'ALL
to read in all variables from the RTO_REGIONS file and then use the names of the variables in that file (RTO_01, RTO_02, etc.) in the ISAM control file, rather than using the custom mapped R01, R02, labels created by your approach. There is some documentation on defining regions in Appendix B of the Users Guide.
I’ll leave it to others to comment on the specifics of shp2cmaq and ISAM, but I have seen the same “Time step not available” warning in DESID_INIT_REGIONS for my own custom region files generated with python. In my case, setting the TFLAG variable of the regions file to an array of zeros with shape (1, n_regions, 2) allowed for CMAQ to read from the regions file.
thanks, Christian and Nash.
I can confirm that the issue was resolved when I forced TFLAG to an array of zeros. I think the issue in shp2cmaq is coming from cmaqsatproc which forces the date to 1970 (screesnshot below), and thus the mask is not seen to be time-independent by ISAM.
For now, I’m using a patched version of the shp2cmaq code which seems to work by making the following two changes:
(1) pass an extra argument to specify that a timeindependent file is wanted
def shp2cmaq(
shppath, attrkey, gdnam, gdpath=None, outpath=None, bcrs='EPSG:4269',
srckey='intersection_area', outformat='NETCDF4_CLASSIC', prefix=None,
overwrite=False, time_independent=True, verbose=1
):
and then
(2) add the following snippet at the end of the method
igf.to_netcdf(outpath, format=outformat)
# add the following
if time_independent:
from netCDF4 import Dataset
import numpy as np
with Dataset(outpath, "r+") as ds:
tflag_len = ds.variables['TFLAG'][0,:].shape[0]
ds.variables['TFLAG'][0,:] = np.array([0,0]*tflag_len).reshape(tflag_len,2)
return outpath